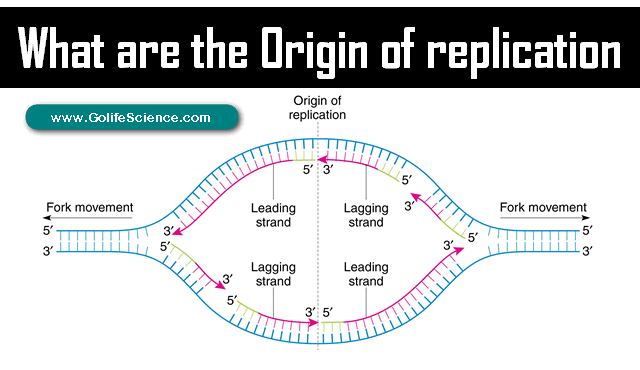
What is the origin of replication? It (also called the replication origin) is a particular sequence in a genome at which replication is initiated.

This can either involve the replication of DNA in living organisms such as prokaryotes and eukaryotes, or that of DNA or RNA in viruses, such as Double-stranded RNA viruses.
What happens first at each origin of replication?
DNA replication may proceed from this point bidirectional or unidirectional.
The specific structure of the replication origin varies somewhat from species to species, but all share some common characteristics such as high AT content.
The replication origin binds the pre-replication complex, a protein complex that recognizes, unwinds and begins to copy DNA.
Origin of Replication in Eukaryotes vs Prokaryotes
The two types of replication origin are:
- Narrow or broad host range
- High- or low-copy number
There are also significant differences between the Replication of origin in Eukaryotes and Prokaryotes :
- Bacteria have a single circular molecule of DNA, and typically only a single replication origin per circular chromosome.
- Archaea have a single circular molecule of DNA and several origins of replication along this circular chromosome.
- Eukaryotes often have multiple origins of replication on each linear chromosome that initiate at different times (replication timing), with up to 100,000 present in a single human cell.
- Having many origins of replication helps to speed the duplication of their (usually) much larger store of genetic material.
- The segment of DNA that is copied starting from each unique replication origin is called a replicon.
Origins of replication are typically assigned names containing ori.
Origin of Replication in Prokaryotes
Bacterial Genome Origins
The genome of E. coli is contained in a single circular DNA molecule of 4.6 x 106 nucleotide pairs. DNA replication begins at a single origin of replication. Replication origin is known as ori-C. In E. coli, the oriC consists of three A-T rich 13-mer repeats and four 9-mer repeats.
10 to 20 monomers of the replication protein dnaA bind to the 9-mer repeats, and the DNA coils around this protein complex forming a protein core.
This coiling stimulates the AT-rich region in the 13-mer sequence to unwind, allowing copies of the helicase dnaB and its co-factor protein dnaC to bind to each strand of the resulting single-stranded DNA.
The dnaB protein forms the basis of the replisome, a complex of enzymes that performs DNA replication.
Bacterial Plasmid Origins
Many bacteria, including E. coli, contain Plasmids that each contains a replication of origin. These are separate from the origins of replication that are used by the bacteria to copy their genome and often function very differently.
For example, the E. coli plasmid pBR322 uses a protein called Rop/Rom to regulate the number of plasmids that are within each bacterial cell.
The most common origin of replication that is used in plasmids for genetic engineering is called pUC. This origin is derived from pBR322 but it contains two mutations.
One single point mutation in the origin itself and another that deletes the Rop/Rom gene.
This removes all the regulatory constraints on the plasmids replication and the bacteria then go from producing 30-40 plasmids per cell with pBR322 up to producing over 500 with pUC.
This allows genetic engineers to produce large quantities of DNA for research purposes. Other origins of replication include pSC101 (derived from Salmonella, around 5 copies per cell), 15A origin (derived from p15A, 10-20 copies per cell) and Bacterial artificial chromosomes (1 copy per cell).

Origin of Replication in Eukaryotes
In eukaryotes, the budding yeast Saccharomyces cerevisiae has the best-characterized replication origins.
These origins were first identified by their ability to support the replication of mini-chromosomes or plasmids, giving rise to the name Autonomously replicating sequences or ARS elements.
Each budding yeast origin consists of a short (~11 bp) essential DNA sequence (called the ARS consensus sequence or ACS) that recruits replication proteins.

In other eukaryotes, including humans, the DNA sequences at the replication origins vary. Despite this sequence variation, all the origins form a base for assembly of a group of proteins known collectively as the pre-replication complex (pre-RC):
- First, the origin DNA is bound by the Origin Recognition Complex (ORC) which, with help from two further protein factors (Cdc6 and Cdt1), load the Mini-Chromosome Maintenance (or MCM) protein complex.
- Once assembled, this complex of proteins indicates that the replication origin is ready for activation. Once the replication origin is activated, the cell’s DNA will be replicated.
In metazoans, the pre-RC formation is inhibited by the protein ‘Geminin’, which binds to and inactivates Cdt1. Regulation of replication, such as this, is important as it prevents the DNA from being replicated more than once each cell cycle.
In humans, an origin of replication has been originally identified near the ‘Lamin B2 gene’ on chromosome 19 and the ORC binding to it has extensively been studied.
Origin of Replication in Viruses
Viruses often have a single Replicative origin.

A variety of proteins have been described as being involved in viral replication. For instance, Polyomaviruses utilize host cell DNA polymerases, which attach to a viral origin of replication if the T antigen is present.
